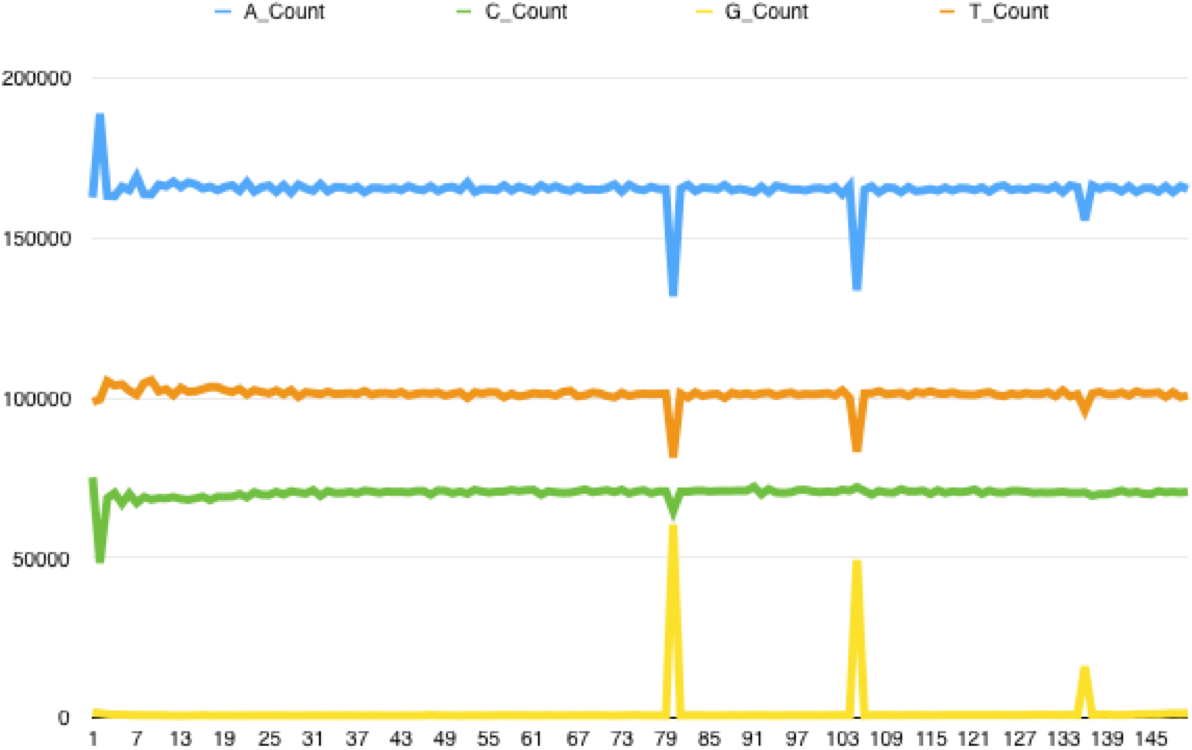
Recently I’ve had a great deal of difficulty with some BS-seq data, which had some bad reads where the quality dropped off randomly in the middle of the sequence – this is unusual as quality issues are typically at the ends of reads. You can see this illustrated below. The poor quality reads seem to cluster within certain tiles on the flowcell.
I asked for help on Biostars, but there were no responses.